|
accurately and psbA-trnH sequence can be the optimal barcode to identify the species of Aloe L. Conclusion: DNA barcoding technology can identify species in Aloe L. while there existed fuzzy identification as to the other three candidate barcodes. The phylogenetic tree constructed by psbA-trnH sequence was capable to identify the six species of Aloe L. Due to some overlaps of intra-and inter-specific variation, there was no obvious barcoding gap in the aspect of ITS2, rbcL, and matK sequences. 2014 Sequencher 5.3 introduces new functionality for RNA-Seq analyses, as well as enhancements to Sequencher Connections. The maximum K2P intra-specific genetic distances were much smaller than the minimum inter-specific genetic distance of all samples in terms of psbA-trnH sequence and thus forming into a prominent barcoding gap. Sequencher 4.1 Sequencher is a DNA sequence assembly and analysis utility 4.3 3 votes Your vote: Latest version: 1 See all Developer: Gene Codes Corporation Review Download Comments (1) Questions & Answers Share 1 / 4 Awards (2) Show all awards Used by 7 people All versions Sequencher 1 (latest) Sequencher 5.3 Sequencher 5. This new release adds features to Sanger, NGS and RNA-Seq analyses. The alignment length and GC contents ranged from 255 to 723 bp and 30.6% to 68.2%. The successful sequencing rate was 100% of all the four candidate barcodes. and compared with the program Sequencher 4.1 (Gene Codes, Ann Arbor, MI). were acquired altogether, with each of ITS2, psbA-trnH, rbcL, and matK regions having 18 sequences respectively. of Sequencing Analysis Program version 4.1 software (Applied Biosystems). Results: Totally 72 sequences of species in Aloe L. The phylogenetic trees were constructed on the basis of Neighbor-Joining method. DNA barcoding gaps were evaluated by the analysis of the intra-and inter-specific divergences. 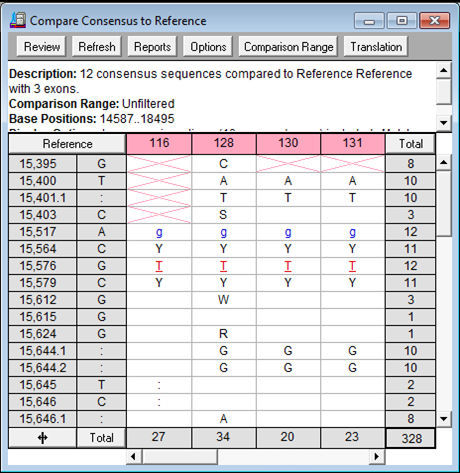  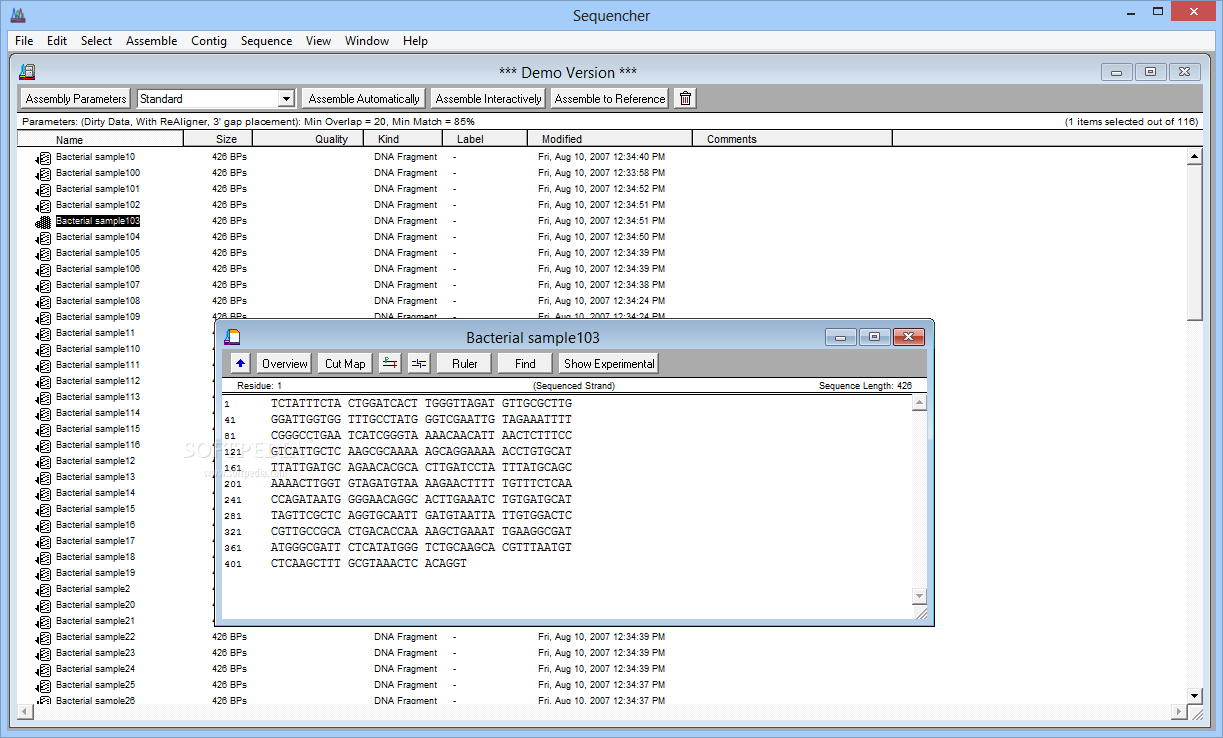
The characteristics of four candidate DNA barcodes were analyzed and the Kimura 2-Parameter (K2P) distances were calculated based on the sequences using MEGA 5.0. The obtained sequences were assembled using Sequencher 4.1.4. was extracted by Plant Genomic DNA Kit, then nuclear ITS2 and the chloroplast psbA-trnH, rbcL, and matK of the samples were amplified using the universal primers and sequenced bidirectionally. Methods: The genomic DNA from 18 samples of six species in Aloe L. ABSTRACT Objective: To screen out a proper DNA barcode for suitable of six species in Aloe L.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
